-
Bioprinting, a cutting-edge technology that merges biology and additive manufacturing, has revolutionized the field of tissue engineering1,2. This innovative approach allows for the precise deposition of biomaterials, cells, and growth factors to fabricate complex, functional tissues and organs3–5. As a result, bioprinting has opened new frontiers in tissue engineering, offering potential solutions for broad applications including disease modeling6,7, drug testing8,9, and regenerative medicine1,10. Common bioprinting methods include inkjet printing11, extrusion-based printing12, laser-induced forward transfer13, and light-based additive manufacturing14–16. Laser- and light-based additive manufacturing has the advantage of high resolution (~1 μm) compared to that of nozzle-based techniques such as inkjet printing and extrusion-based printing (~100 μm)17. Bioprinted implants typically involve a surgical intervention for the implantation18,19 or for direct in-situ biofabrication at the exposed site20,21, which poses inherent challenges and risks.
To address these limitations, non-invasive and minimally invasive bioprinting has emerged as a powerful solution by offering the possibility of creating functional biological constructs bypassing invasive surgical procedures22–25. Light-based additive manufacturing, which employs light to solidify resins without the need for direct material deposition, is particularly well positioned compared to other minimally invasive bioprinting techniques thanks to light and its possibility of delivering energy through tissues. More precisely, the light energy is sent through the tissue to initiate photopolymerization of the injected bio-ink and transform it into desired structures. Light transport in biological tissues is determined by their absorption and scattering properties. Ultra-violet (UV) or blue light, which is commonly used in photopolymerization, shows poor tissue penetration depth and is not favorable for non-invasive bioprinting. The near-infrared (NIR) window with a wavelength ranging from 650 to 1350 nm, offers deeper penetration into biological tissues with less significant attenuation because of its longer wavelength (less scattering) and the lack of absorption from biological molecules26. Therefore, NIR light is well-suited for in vivo imaging27–30 and therapeutic applications30–32 that require light to reach target areas deep within the body. It can induce photopolymerization via two-photon absorption33 or upconverting process34 and has already been demonstrated in non-invasive additive manufacturing22–24.
Although NIR light is transmitted more efficiently through tissues, scattering still scrambles the propagating light field to form complex speckle patterns, preventing focusing the light to a tiny spot for a well-confined delivery of light energy. This greatly impacts the resolution (from 1 μm in the absence of tissue to tens or hundreds μm depending on tissue properties and thickness) and the fidelity of non-invasive printing22,24. In the field of optical imaging, several strategies based on wavefront shaping have been developed to manipulate the light and refocus it through scattering media to a diffraction-limited spot35–41. These techniques utilize feedback signals obtained behind the scattering media to spatially modulate the input light in phase and amplitude. For non-invasive light focusing, fluorescent or acoustics signals emanating inside or behind the scattering medium can be measured from the same side as the light delivery38,39,41. However, these techniques only provide the wavefront information of one target location at a time, and it is time-consuming to refocus at each voxel to be printed. Fortunately, the scattered optical field preserves a certain degree of correlation, which is commonly referred to as the optical memory effect42,43. When an input wavefront reaching a scattering medium is shifted (or tilted) within a certain distance (or angle), the output wavefront propagating through the medium is equally shifted (or tilted). In thick biological media, where scattering is anisotropic (anisotropic factor g usually ranges from 0.9 to 0.9826), the range of tilt/tilt memory effect becomes minimal (50-μm-thick tissues around 3-8 mrad44) but strong shift/shift correlations are still observed43. In this way, the focal spot can be shifted through the scattering medium before it becomes too dim so that the next focusing optimization can be generated in a time-efficient manner45. The scattering effect of the tissue is thus corrected during the printing using sparse focusing, which significantly speeds up the printing as compared with optimizing at every subsequent spot.
In this study, we develop a micro-meter resolution additive manufacturing technique through a highly scattering medium assisted by upconversion nanoparticles (UCNPs). As the UCNP generates fluorescence of different wavelengths under the illumination of NIR light, it acts not only as a secondary UV source for photopolymerization but can also be used as a guide star for the feedback loop to refocus light through the scattering media. Then, the focal spot is scanned through the scattering medium using sparse focusing. Based on this technique, we are able to print high-resolution (2 μm) structures through a holographic diffuser and a chicken tissue of thickness 300 μm. These results demonstrate high-resolution additive manufacturing through strongly scattering media and suggest potential applications in non-invasive biomedicine.
-
We designed our non-invasive additive manufacturing system based on wavefront shaping, as illustrated in Fig. 1. A NIR beam at 976 nm is first modulated in amplitude by a digital micro-mirror device (DMD) and directed through a scattering medium (holographic diffuser or chicken tissue) into the resin. The resin contains hydrogel monomers of gelatin methacryloyl (gelMA) and UCNPs coated with the UV light photoinitiator lithium phenyl-2,4,6-trimethylbenzoyl-phosphinate (LAP). The synthesized UCNPs are highly crystalline and show hexagonal morphology, with an average particle size of ~10 nm (Fig. S1). UCNPs (positive) are coated with LAP (negative) through electrostatic interaction. This coating is verified by Fourier transform infrared (FTIR) and Zeta potential analysis (Fig. S2). UCNPs emit UV and visible fluorescence under the illumination of 976 nm light. The visible fluorescence (440 nm < λ < 550 nm), which is not absorbed by LAP, is back-scattered by the scattering medium and epi-detected by a single-photon avalanche diode (SPAD), providing the feedback signal for the optimization of the spatial light modulation (binary DMD pattern). Then, the optimized DMD pattern is displayed and refocuses the NIR light through the scattering medium to a diffraction-limited spot within the resin, which is shifted together with the scattering medium to induce photopolymerization along the designed path. After a lateral shift of the sample determined by the size of the memory effect (here around 5 μm), the DMD pattern is re-optimized to focus light again and this scanning process is repeated until the printed part is complete. The resulting spatial distribution of the NIR light at the focal plane is inspected by a camera placed on the distal side of the sample, for observation purposes only.
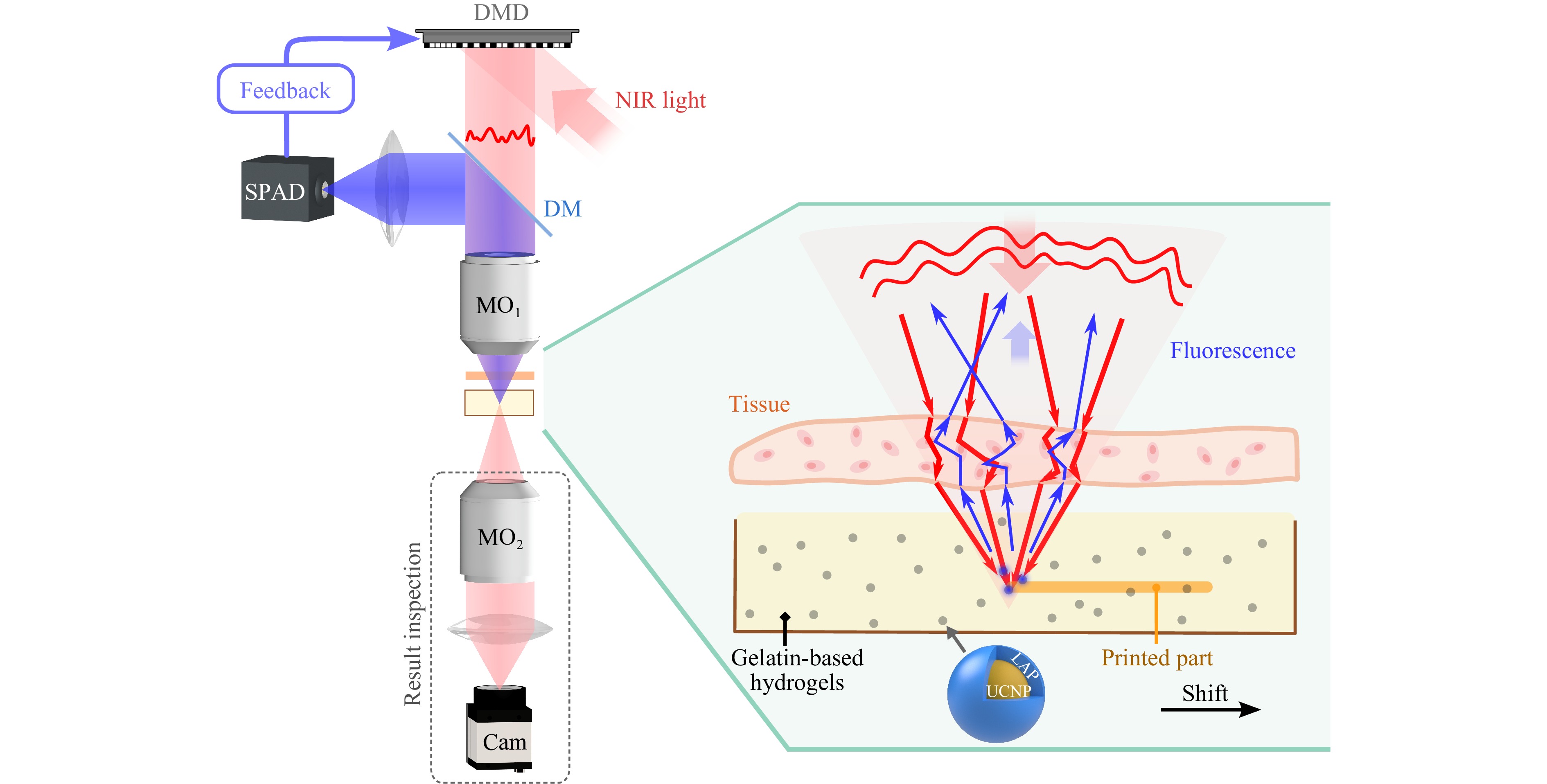
Fig. 1 Schematic figure of high-resolution non-invasive additive manufacturing using UCNPs that are coated with the photoinitiator LAP. The NIR beam is modulated by the DMD to compensate for the scattering and focus the light through the tissue down to a diffraction-limited spot. The resin contains UCNPs that convert NIR light to UV and visible fluorescence, acting as the secondary UV source for inducing the photopolymerization of the hydrogels and as a guide star for wavefront shaping. By laterally shifting the sample across the print geometry, micro-meter resolution features can be printed through the tissue. DM: dichroic mirror. MO: microscope objective. Cam: camera.
The focusing process using the upconverted fluorescence as feedback is shown in Fig. 2. UCNPs convert NIR light to UV and visible light (Fig. 2a). The latter conveys information about the NIR speckle pattern within the resin. Fig. 2b shows the emission spectrum of UCNPs illuminated by 976 nm light and the absorption spectrum of the photoinitiator LAP. The emission peaks at 350 and 360 nm fall within the absorption band of LAP, suggesting that it is mainly absorbed by the LAP coating and contributes to photopolymerization. The rest of the fluorescence can be partially detected in reflection thanks to its isotropic emission. Upconverted fluorescence in the wavelength range of 440 nm < λ < 550 nm is experimentally chosen as the feedback for the following consideration. The upconversion process to a high-energy photon involves multi-photon absorption, resulting in a nonlinear luminescence process. Each fluorescence peak corresponds to a certain nonlinearity parameter n, which can be understood as the number of NIR photons absorbed required to emit a photon of higher energy than the incident NIR photons. Due to the saturation of the excited energy states, the nonlinearity is experimentally experienced only at low light intensity. In the focusing process, signals generated from a high nonlinearity conversion are preferred because of their faster converging speed38,46,47. However, this signal only occurs at low intensity and shorter wavelength48 and thus there is a balance between non-linearity and signal intensity since a higher photon count enables a faster collection and speeds up the focusing process. Upconverted fluorescence in this wavelength range is chosen because it covers most of the photons from visible emission and preserves a high average non-linearity. The total fluorescence is measured by the SPAD at different NIR intensities and plotted in the log scale (Fig. 2c). The NIR intensities are calculated by the laser power measured before the illumination objective divided by the beam size at the focal plane, and the transmission of this objective (~67% at 976 nm) is not taken into account. According to the definition, the slope of the curve in the log scale represents the nonlinear parameter n. The total fluorescent signal collected displays a slope of 2.4 at a lower intensity and a decreased nonlinearity at a higher intensity.
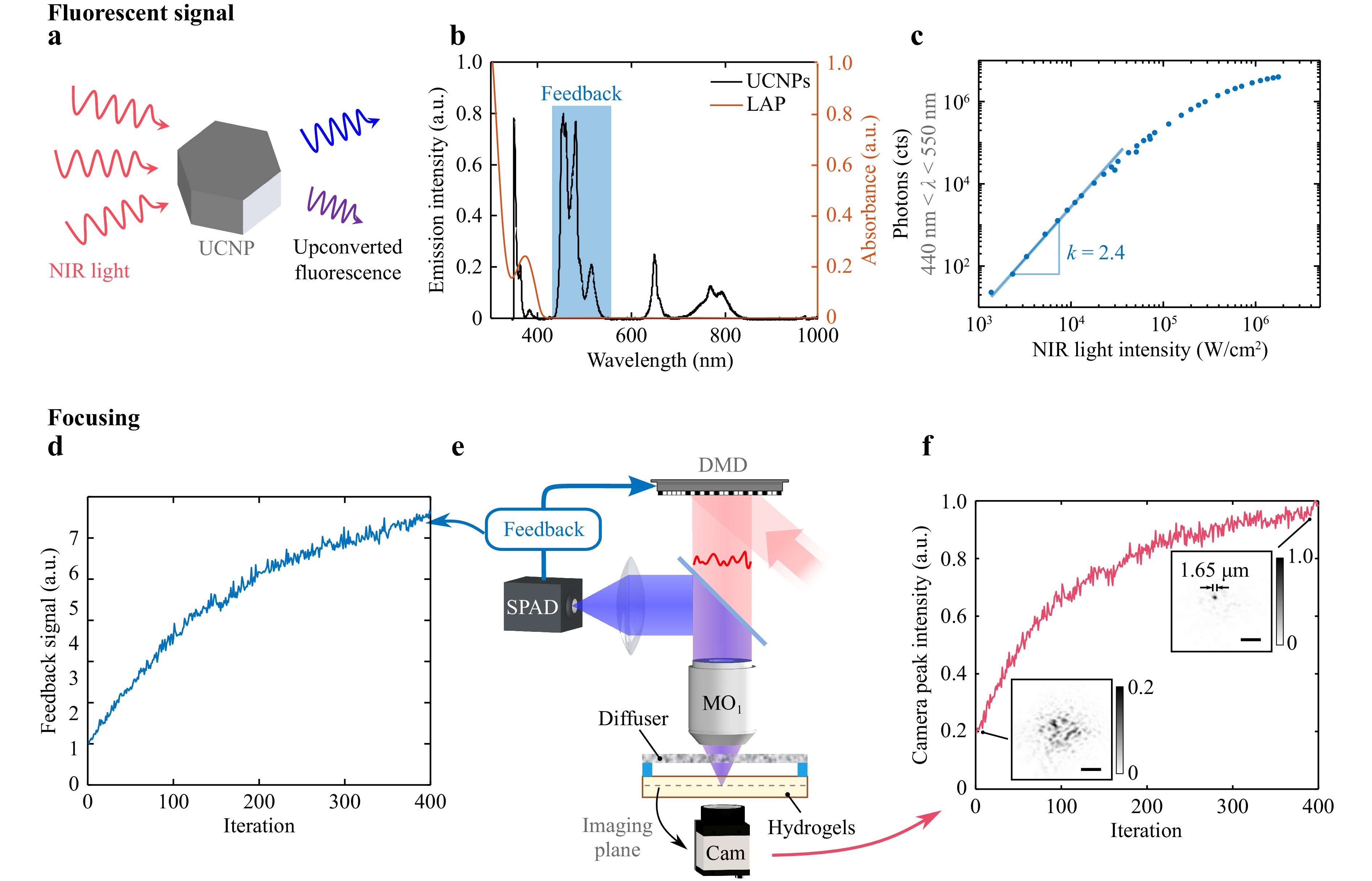
Fig. 2 Focusing process based on the nonlinear fluorescent feedback. a Schematic diagram of the upconverting process. b Emission spectrum of UCNPs (black) under 976 nm light illumination and absorption spectrum of the photoinitiator LAP (orange). The wavelength band highlighted in blue represents the range of upconverted fluorescence collected as the feedback. (a.u.= arbitrary units). c Fluorescent feedback versus NIR light intensity in the log scale. The slope represents the nonlinearity parameter n. d Power-corrected feedback signal during the iterative optimization. e Experimental setup for non-invasive focusing. A holographic diffuser is used in this experiment. f Peak intensity of the NIR patterns inspected by the camera during the optimization. Insets show the NIR patterns at the first and the final iteration. Full width at half maximum (FWHM) of the focal spot, 1.65 μm. Scale bars, 10 μm.
Because of the low upconversion efficiency of UCNPs, the total fluorescence is detected by a single-pixel detector to ensure a good signal-to-noise ratio (Fig. 2e), at the expense of a loss of spatial information of the speckle. As already demonstrated38,46,49, optimizing a nonlinear spatially integrated signal enables blind focusing behind a scattering layer through iterative optimization. By maximizing the total fluorescent signal, the light tends to redistribute the energy to one single spot rather than over several grains of a speckle thanks to the nonlinear fluorescence behavior at the chosen NIR intensity. Note that, we have no control over the position of the focal spot, which can be at any hot spot of the speckle illuminating the resin. In most of the previous research works38,46,47,49, the light is modulated in phase with a liquid crystal-based spatial light modulator (LC-SLM), which does not change the light power after the modulation. In this work, however, a DMD is implemented because of its faster operation (~20 kHz) compared to that of LC-SLM (~60 Hz). Therefore, the light is modulated only in amplitude, resulting in pattern-dependent output power. The fluorescence signal is also dependent on the number of pixels on the DMD with the “ON” state, which does not necessarily result in a focal spot. Therefore, the fitness function $ f\left(x\right) $ for a DMD pattern Z is calculated as:
$$ f\left(\mathit{Z}\right)=\frac{{P}_{fluo}}{{\mathit{I}}_{DMD}\odot \boldsymbol{Z}} $$ (1) Pfluo is the total fluorescent signal resulting from this DMD pattern. $ {\mathit{I}}_{DMD}\odot \boldsymbol{Z} $ is the element-wise product of NIR light distribution on DMD (Fig. S3) and the DMD pattern Z, which gives the light power of this pattern before the illumination objective. By maximizing this fitness function, the iterative algorithm tries to find the DMD pattern that excites more fluorescence per NIR light power, which compensates for the effect of amplitude modulation. In the iterative optimization, we adopted separable natural evolution strategies50 (SNES) to increase the converging speed of the global search and shorten the optimization time. Multiple pixels are encoded with a number between 0 and 1 (Fig. S4a) to eliminate the drastic change between pixels in the binary amplitude modulation50. The focusing process is operated at low NIR power (average intensity of ~3×103 W/cm2): the nonlinearity parameter n is large, resulting in a faster converging speed; the light dose is much lower than the photopolymerization threshold so that it does not induce photopolymerization. The camera placed on the other side of the sample is only used for imaging the NIR pattern. Because the focal spot can converge at any position of the resin volume illuminated by the speckle, the resin is contained in a rectangular capillary with an inner thickness of 20 μm to limit the position of the focal spot along the optical axis, making it easier for the alignment of the imaging system (see Section S5, Supplementary Information). By maximizing the feedback signal mentioned above (Fig. 2d), the peak intensity on the image of the NIR pattern increases with the iteration (Fig. 2f), leading to only one sharp spot behind the diffuser. The diffraction limit of this system is determined by the numerical aperture (NA). In this study, we use an NA 0.40 objective to focus the light after DMD projection. The NA of this system is ~0.40 after passing through the scattering layer because both the holographic diffuser and the tissue are anisotropic scattering media. Therefore, the theoretical diffraction limit is d = λ/NA = 1.22 μm. FWHM of the focal spot measured from 5 different samples is 1.65 ± 0.13 μm. It is slightly larger than the theoretical diffraction limit, which is mainly because of the following reasons. First, the modulated beam is slightly smaller than the back aperture of the objective in order to use the full pattern of the DMD. Therefore, it results in a smaller NA. Secondly, the optimization iteration does not fully converge before being stopped, which is a balance between the spatial resolution and the optimization time.
After forming a sharp spot behind the diffuser, the optical memory effect can be measured by shifting the sample laterally (Fig. 3a). The capillary containing the resin is fixed onto the diffuser by a spacer (1 mm). Shifting the diffuser together with the resin is equivalent to shifting the beam except that the focal spot will remain at the same position on the image captured by the camera, making the inspection easier. Fig. 3b shows that the peak intensity of the focal spot decreases with the distance Δx from the original position. For this holographic diffuser, the full width at half-maximum (FWHM) of the memory effect range is 16 μm.
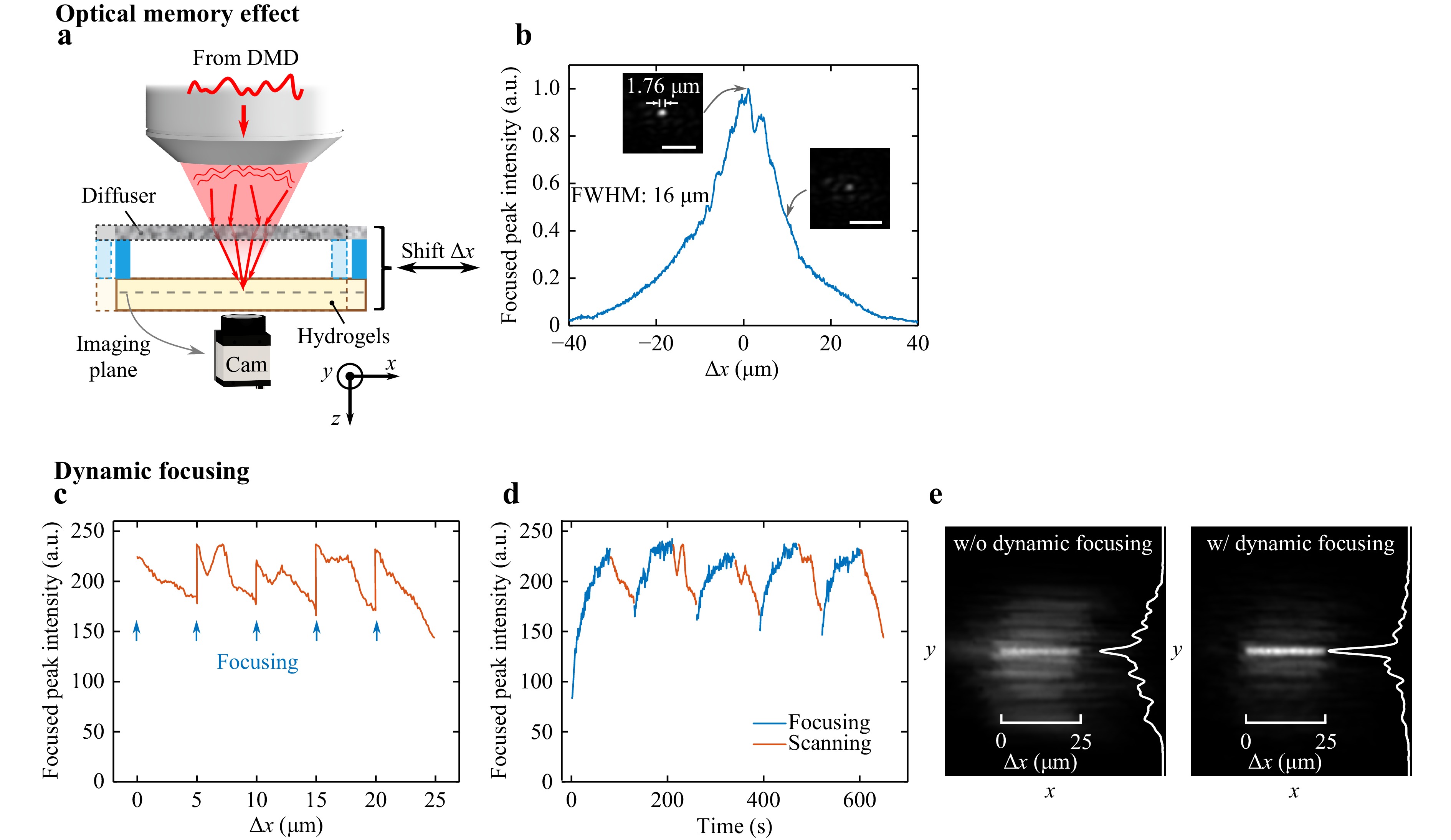
Fig. 3 Dynamic focusing and scanning based on the optical memory effect. a Schematic figure of the setup showing the lateral shifting of the sample. b Focused peak intensity with the DMD pattern from the initial focusing versus the shifted distance for the holographic diffuser. Insets show the focal spot at Δx = 0 and Δx = 10 μm. FWHM of the focal spot, 1.76 μm. Scale bars, 10 μm. Focused peak intensity versus the shifted distance c and time d during the dynamic focusing. The scanning process results in a decrease in peak intensity at the focal spot. The focusing process is repeated every 5 μm to maintain a sharp and intense focus. e NIR light dose without and with dynamic focusing after a shifted distance of 25 μm. The curve on the right-hand side of each image shows the NIR intensity profile along Δx = 12.5 μm.
Thanks to the memory effect, we are able to scan the focal spot within the field of view without changing the DMD pattern at each printed voxel. However, the spot intensity decreases with the shifting distance, as well as the contrast (Fig. S6), and a high-resolution structure can only be fabricated within a small area around the initial focusing position. In addition, the light dose is not uniform, resulting in different degrees of polymerization across the structure. To maintain a similar light dose at each voxel to be printed, we adopt dynamic focusing. The focused spot is shifted across the diffuser over 5 μm before the re-focusing process starts again (Fig. 3c). The optimized pattern from the previous focusing process serves as the initial pattern for the new optimization, which greatly increases the converging speed compared to the optimization from scratch45. The peak intensity of the first iteration of the focusing is lower than that of the ending position of the last scanning process (Fig. 3d) because random deviations are introduced to the initial pattern in order to find the global maximum. Because of the low fluorescent signal collected, the speed of the focusing process is limited to the integrating time of the SPAD for each display. A DMD framerate of 300 Hz is used to ensure that the signal has enough signal-to-noise ratio to reflect the information of the speckle. The focusing time is approximately 90 s (limited by the fluorescence collection) and the scanning time is approximately 50 s (limited by the required dose to solidify the resin) for each 5 μm of lateral shift. The focal spot maintains a relatively stable intensity during the whole process. Fig. 3e shows the NIR light dose distribution on the focal plane with and without dynamic focusing. The experiment without dynamic focusing is conducted by laterally shifting the speckle over the same total distance (25 μm). The light dose distribution is calculated by summing up the NIR speckle patterns during the scanning process according to the shifting distance. The profile at Δx = 12.5 μm is plotted on the right-hand side of each image to show the contrast. With dynamic focusing, we are able to create a dose distribution in the shape of a sharp line with a uniform intensity.
Invasive printing is first conducted in order to explore the possibilities of this technique in tissues without the limitation of the low fluorescent signal after back-scattering. It is demonstrated using both a holographic diffuser and a slice of 300-μm-thick chicken breast. We call this invasive printing because the SPAD detector is placed on the distal side of the sample (Fig. 4a). It is worth stressing that this is the only case in this study, in which the feedback signal is collected in a transmission-based configuration. As the fluorescence is directly collected from the emission site, and thus not experiencing strong loss due to back-scattering, the integration time of the SPAD can be significantly decreased, reducing the optimization time down to 20 s (the DMD displays patterns at 1 kHz during the iterative optimization). The framerate is limited by the long rise time (~0.2 ms) and decay time (~0.3 ms) of the upconverted luminescence of our UCNPs (see Section S7, Supplementary Information). With a faster light modulation, multiple illumination patterns might contribute to the measured fluorescent signal, resulting in inaccurate feedback. The optical memory effect of the chicken tissue has a FWHM of 8 μm, smaller than that of the diffuser (Fig. 4b). Fig. 4c and e show qualitatively the impact of scattering on the text readability placed underneath for the holographic diffuser and the chicken breast layer respectively. Letters “EPFL” are printed to verify the capability of printing length scale several times the optical memory lateral shift. During the printing, the focusing process is performed at low NIR power (to benefit from the non-linearity of the fluorescence signal), and lateral scanning is performed at high NIR power to ensure that the light dose at the focal spot surpasses the polymerization threshold. Fig. 4d and f show the printed structures imaged by a differential phase contrast (DPC) microscope51, which is used in this study because of the low refractive index mismatch between the polymerized and unpolymerized hydrogels (see Section S8, Supplementary Information). The bright and dark edges in the DPC image represent the distribution of phase change (refractive index mismatch) and its contrast (bright minus dark intensity) is positively correlated with the strength of phase change, hence the degree of photopolymerization in this study. The initial focal spot of each letter (top-left corner) is optimized from scratch (a speckle pattern) and the rest is completed with dynamic optimization (from a dim focal spot). The printed structure through the holographic diffuser is relatively uniform, matching the result of the light dose (see the supplementary video). Looking in detail, the line at the bottom of the letter “E” is detached from the rest of the letter at the bottom-left corner. This is because we have no control over the position of the focal spot (global maximum), the position of which happens to switch at this position. For the printed part through the chicken tissue, over-polymerization can be seen in the letter “P” and “F”, while optimization did not converge completely when printing the letter “E”. It is very likely that the non-uniformity of muscle fibers in the chicken breast results in different scattering properties across the tissue. The tissue structure above the letter “E” is probably more scattering and requires longer focusing time while the tissue above letters “P” and “F” is less scattering, resulting in a brighter focal spot and a higher degree of polymerization. To improve the fidelity in practical applications, the criteria for stopping the optimization process should be based on the converging speed of the feedback signal, ensuring that a focal spot is formed in each region. As the optimized feedback signal reflects the intensity of the focal spot, the laser power for printing can be adjusted in real-time to achieve a similar degree of polymerization across the whole structure.
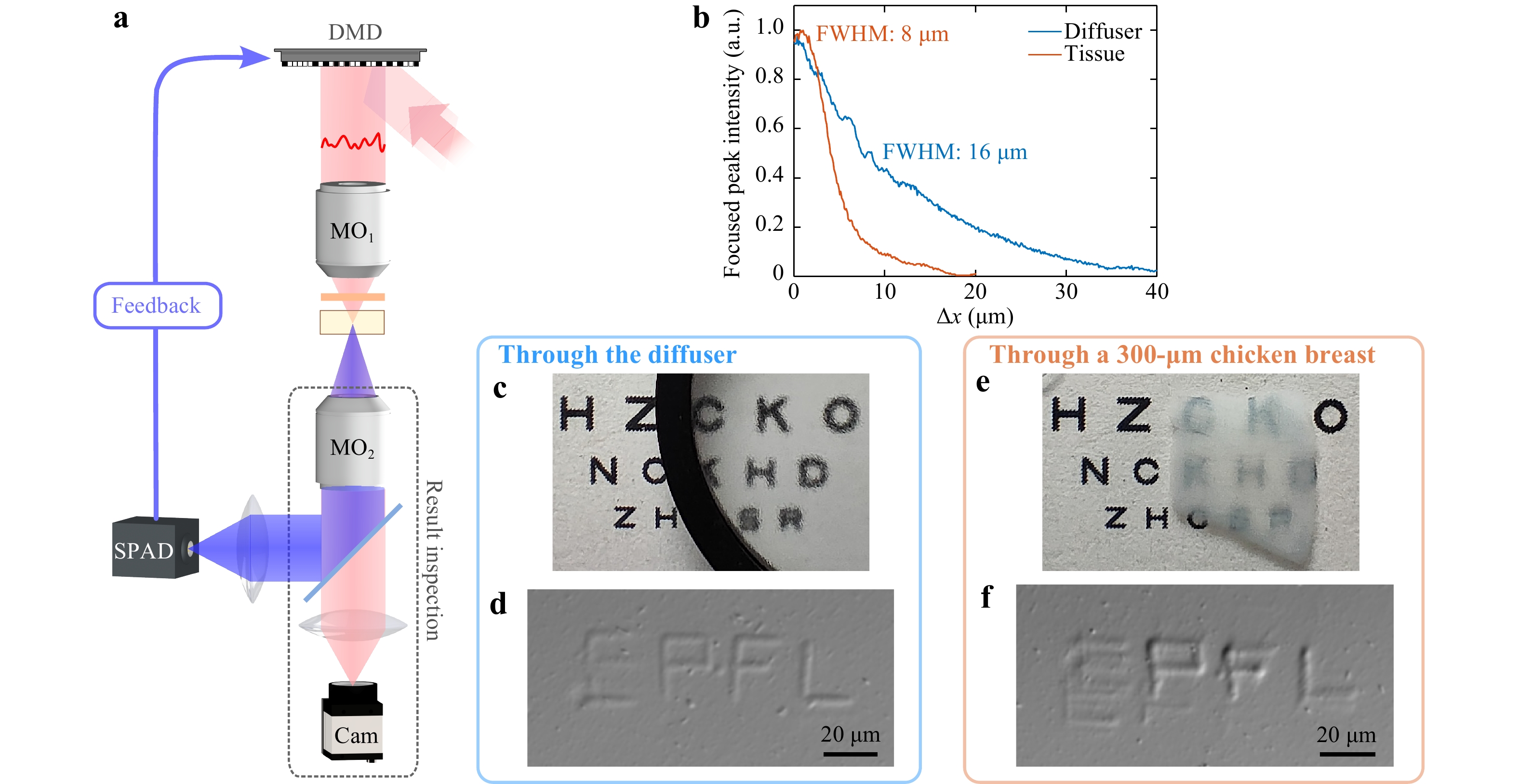
Fig. 4 Invasive printing. a Experimental setup for invasive printing. b Focused peak intensity versus shifted distance for the holographic diffuser (blue) and for the chicken tissue (orange). Photographs of a target placed below the diffuser c and the tissue e. These scattering media prevent visually differentiating the letters of the bottom line. DPC images of printed “EPFL” through the diffuser d and the tissue f.
Non-invasive printing is then demonstrated through the diffuser by placing the SPAD detector at the same side as the light delivery (Fig. 5a). The chicken tissue is not tested in this configuration: the tissue exhibits excessive scattering at shorter wavelengths, significantly decreasing the amount of epi-detected (reflection mode) fluorescence which translates into a too-low optimization speed for printing. Based on dynamic focusing, we are able to print fine structures within the speckle (Fig. 5b). Without dynamic focusing (DMD acts only as a mirror), only hot spots in the speckle are printed. By shifting the sample laterally, lines of different contrast and lengths are photopolymerized within the areas highlighted by dash lines (Fig. 5c). As the intensity of the speckle grains decreases at the edge of the speckle, there is no sharp boundary between the polymerized and unpolymerized area, deteriorating the printing fidelity and the printing resolution. The sample is also laterally shifted along the path of the letter “E”. As expected, no resolvable structure can be printed inside the speckle size without dynamic focusing (Fig. 5d). In contrast, with dynamic focusing, sharp and uniform lines with a feature size of 1.78 ± 0.39 μm (Fig. S8b) can be printed and the minimum resolvable distance that we obtained is 2 μm (Fig. 5e). Feature size is characterized at 5 different regions of the structure. Fig. 5f shows a clearly printed letter “E” which is even smaller than the speckle size. Although we have no precise control over the absolute position of the printing (it may start at any place inside the speckle), the relative position of the structures is controlled accurately.
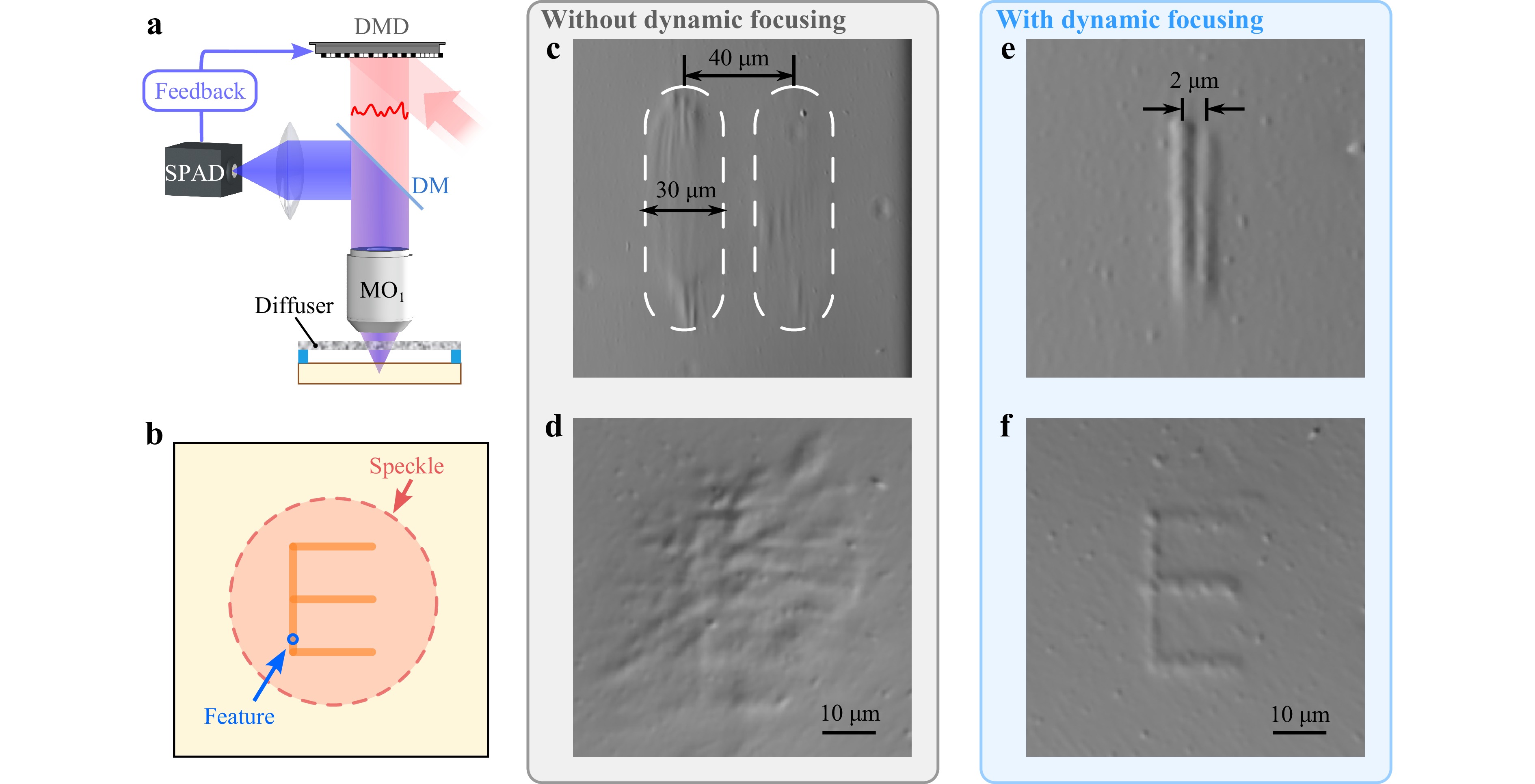
Fig. 5 Non-invasive printing. a Experimental setup for non-invasive printing. b Schematic diagram of a comparison of the speckle size and the printed feature size. Fine structures can be printed accurately inside the speckle. c DPC image of two “lines” printed without dynamic focusing and at a distance of 40 μm. The polymerized area is roughly highlighted by white dashed lines because no clear boundary can be observed. d DPC image of a polymerized part following the path of the letter “E” without dynamic focusing. e DPC image of two printed lines with a center-to-center distance of 2 μm using dynamic focusing. f DPC image of a printed letter “E” using dynamic focusing.
-
Focusing light through strongly scattering media had long been considered impossible until the recent progress in the field of wavefront shaping. Driven by the ever-growing need for deep tissue in vivo imaging30,52, non-invasive imaging has been achieved through various techniques such as blind focusing with nonlinear signals38,49, fluorescence-based transmission matrix41,53, and acoustic manipulation40. In this work, we use blind focusing which utilizes nonlinear signals to cope with the issue of low feedback signal. This has been demonstrated with two-photon imaging38 and three-photon imaging46,49 and should be also available for other nonlinear processes54. For linear signals that have enough signal intensities, non-invasive focusing and imaging can be achieved by collecting the back-scattered fluorescent patterns41,53. Nano-focusing devices55,56 such as metalens on the end face of a single-mode fiber have also been proposed for imaging applications. The techniques readily available in the wavefront shaping might inspire the development of new bioprinting methods against the turbid nature of the biological tissue.
In this work, we make use of an iterative wavefront method to enable non-invasive additive manufacturing through a scattering layer. The light propagating through the scattering medium produces complicated speckle patterns that locally excite UCNPs. Because of the nonlinear upconverted fluorescence as the feedback, a sharp focus can be formed in the resin at a fast optimization speed even with a low signal level. Dynamic focusing is conducted to ensure a uniform light dose to solidify voxels and combined with the memory effect, the printing time is optimized. A printing scale 5 times the memory effect size is demonstrated. Successful printing through a diffuser and a 300-μm chicken tissue proves the feasibility of this technique. In the non-invasive configuration, we show that a micro-meter resolution structure can be printed within a speckle pattern that is more than 20 times larger than the feature size. To the best of our knowledge, this study is the first report on high-resolution non-invasive printing through a highly scattering medium, which pushes the boundaries of noninvasive printing to micro-meter resolution.
There is much work to be done before the proposed technique can become a tool for non-invasive in vivo printing. Biological tissues are dynamic scattering media40,57,58 and therefore the time required to focus light in our approach cannot exceed the ms range. The optimization speed in the non-invasive focusing is limited by the signal intensity, which can be increased by using a larger and more sensitive photon detector such as photomultiplier tubes and more importantly improving the upconversion efficiency of the nanoparticles59. SPAD arrays might also be used to gather the spatial distribution of the fluorescence. Noise-tolerant algorithms with relatively fast converging speed are preferred and deep learning might also be implemented for efficient focusing through living tissues60. In terms of biocompatibility, the hydrogels selected in this study are biocompatible and have been widely used in various biomedical applications61. Cell viability experiment of UCNPs also indicates their good cellular compatibility (see Section S9, Supplementary Information).
Real applications also pose requirements in the spatial domain. The nature of blind focusing in this technique denies the possibility of pre-determining the absolute position of the voxel. In this work, we observe that the global maximum will remain at a hot spot for approximately 25 μm of shifting in the lateral direction before switching to another hot spot within the illuminated resin. This determines the printing area of one object (see Section S10, Supplementary Information). The optical memory effect also exists in the axial direction62, indicating the possibility of true 3D printing. As for a larger volume, recent progress in imaging beyond the memory effect53 might help to push this boundary. For applications that require an accurate absolute position of the printed voxels such as connecting the neuron fiber, feedback with spatial information is necessary.
In summary, we have presented a non-invasive additive manufacturing technique to print a hundred-micron structure size through a strongly scattering medium at micro-meter resolution based on the fluorescent feedback from the printing system. Thanks to the nonlinear upconverted fluorescence and the optical memory effect, sub-speckle printing is demonstrated on a 25-μm size print with a printing resolution of around 2 μm. This technique provides a promising route toward high-resolution non-invasive bioprinting and shines light on the development of new techniques for minimally invasive and non-invasive biomedicine.
-
Chemicals used in this experiment were purchased from Merck & Co (Sigma-Aldrich) and the synthesis was carried out in a bifold Schlenk line under the flow of argon gas.
In a typical synthesis, thulium (III) acetate hydrate (0.004 mmol) was reacted with oleic acid (6 mL) and 1-octadecene (15 mL) at 140 °C under partial vacuum having argon atmosphere for 90 min in a 100 mL 3-neck Schlenk flask to prepare oleate solution. Once the reaction was complete, first, ytterbium (III) acetate hydrate (0.240 mmol) was reacted with the above oleate solution for 90 min and afterward, yttrium (III) acetate hydrate (0.556 mmol) was reacted with this oleate solution at 140 °C. This mixed oleate solution, thus obtained, was cooled down to 50 °C. To this, methanol solution (10 ml) of ammonium fluoride (3.2 mmol) and sodium hydroxide (2 mmol) was added dropwise and stirred for 30 min. Methanol was completely removed under partial vacuum and the reaction mixture was further heated to 300 °C (~10 °C/min) under argon and maintained for 60 min. The reaction was frozen by the addition of cold ethanol and the nanoparticles were collected by centrifugation, redispersed in cyclohexane. This process was repeated thrice, before the product being used as core UCNP (0.5 mol% Tm3+, 30 mol% Yb3+ doped) in the next step.
Similarly, in the second step, yttrium (III) acetate hydrate (0.8 mmol), oleic acid (6 ml) and 1-octadecene (15 ml), methanol solution (10 ml) of ammonium fluoride (4 mmol) and sodium hydroxide (2.5 mmol) were used to prepare the NaYF4 shell precursor.
-
Layer-by-layer successive epitaxial shell growth of NaYF4 was achieved on NaYF4:Yb/Tm core UCNPs. Core UCNPs were added to 1-octadecene (5 mL) in a 3-neck Schlenk flask and heated to 300 °C in an argon atmosphere. To this, shell precursor solution was injected @ 5 μL/sec using a Nemesys syringe pump system. The ripening was done at 300 °C for 30 min. After ripening, the reaction was frozen and the core-shell UCNPs were precipitated and washed as outlined for core UCNPs and finally dispersed in hexane (5 mL). These dispersed particles were neutralized using 2M HCl to get the ligand-free core-shell UCNPs.
-
10 g of gelatin (Sigma-Aldrich) was dissolved in 100 mL of phosphate-buffered saline. Then 8 mL of methacrylic anhydride (Sigma-Aldrich) was added dropwise (0.5 mL/min) and the mixture was left under stirring at 50 °C for 3 hours, followed by removal of unreacted anhydride by centrifugation and dialysis against distilled water. GelMA was obtained after lyophilization.
-
Lithium phenyl-2,4,6-trimethylbenzoyl-phosphinate (LAP) (Sigma-Aldrich) was dissolved in water at a concentration of 20 mg/mL. 10 μL of UCNP aqueous solution (100 mg/mL) was mixed with 50 μL of LAP solution and sonicated for 30 min. 15 mg of gelMA was dissolved in the mixture and 40 μL water was added to form a final concentration of 10 mg/mL UCNP, 10 mg/mL LAP and 15 wt% gelMA. The resin was stored at 4 °C until further use.
A uniform distribution of UCNPs in the resin is a critical factor in focusing and printing. If they are not uniformly distributed or even appear in the form of clusters, the optimized focal spot will always be located at the concentrated region. The distribution of UCNPs in the resin was checked by the fluorescent profile of a collimated 976 nm beam at the transverse plane (Fig. S13). The resin preparation process was optimized to ensure a uniform fluorescent profile.
-
Transmission electron microscopy images were acquired on a Tecnai Osiris electron microscope, with an accelerating voltage of 200kV. FTIR spectra were recorded by making KBr pellets of the power samples and measuring them on a Spectrum 3 spectrophotometer (PerkinElmer). Zeta potential was recorded by Nano ZS (Malvern) using dynamic light scattering. The UV-Vis spectrum of LAP was recorded on a Lambda 365 UV/Vis spectrophotometer. The upconverted fluorescence emission spectrum was recorded on a setup as previously reported48. DPC images were recorded on a microscope as previously reported51.
-
A continuous-wave laser at 976 nm (900 mW, BL976-PAG900, Thorlabs) with a Polarization-Maintaining (PM) optical fiber is collimated by a lens (F810APC-1064, Thorlabs). After modulated by the DMD (V-650L, Vialux), the NIR light is directed through the objective MO1 (M Plan Apo NIR 20X, NA 0.40, Mitutoyo) to excite the UCNPs in the resin placed below the scattering medium. The DMD is imaged to the back focal plane of MO1. The scattering medium is a holographic diffuser (Newport 5°) or a slice of fixed chicken breast. The upconverted fluorescence is back-scattered by the medium, collected by MO1 and a lens (f = 15 mm), and detected by a SPAD (PDM-50-CTD, Micro Photon Devices). We use two longpass dichroic mirrors (DMLP550R, Thorlabs, FF699-FDi01-t1-25x36, Semrock) and two shortpass filters (FESH0600, Thorlabs and FF01-720/SP-25, Semrock) to narrow the spectral bandwidth. The NIR speckle patterns are imaged in transmission via MO2 (LIO-40X, NA 0.65, Newport) and a lens (f = 150 mm) onto the Cam (acA2040-55um, Basler). This part of the setup is only used for monitoring the NIR speckles. In the invasive configuration, the fluorescence is collected in transmission through MO2 and a lens (AC254-030-A, Thorlabs), reflected by a longpass dichroic mirror (FF552-Di02-25x36, Semrock), filtered by two shortpass filters (FESH0600, Thorlabs and FF01-720/SP-25, Semrock), and detected by the SPAD.
-
The optimization was done with the SNES algorithm50 (see algorithm flow chart in Fig. S11, Supplementary information). The initial Gaussian parameter μ is a random array from [0,1) with a length of segment number and σ is an array of the same length filled with ones. During optimization, each segment of DMD is parameterized by μ and σ. Npop grayscale patterns are generated according to $ \mathrm{\mu }+\mathrm{\sigma }{s}_{n} $, where $ {s}_{n}(n=1, 2, \dots ,{N}_{pop}) $ is an array with a length of segment number and its elements follow the standard normal distribution. Then these grayscale patterns are converted to binary patterns via a multi-pixel encoding method50 and displayed by DMD. SPAD is synchronized with DMD to collect total fluorescent signals of each binary pattern. After that, all the patterns are sorted according to their power-corrected feedback signals in increasing order and multiplied with weights $ {u}_{n} $. The weights for the first Npop/2 patterns are set to 0 and the rest are set as an arithmetic sequence with a sum of 1. The natural gradients for μ and σ are calculated by Eq. 2.
$$ \left\{\begin{array}{l}{\nabla }_{\mu }J={\displaystyle\sum }_{n=1}^{{N}_{pop}}{u}_{n}\cdot {s}_{n}\\ {\nabla }_{\sigma }J={\displaystyle\sum }_{n=1}^{{N}_{pop}}{u}_{n}\cdot ({s}_{n}^{2}-1)\end{array}\right. $$ (2) The updated μ and σ are calculated by Eq. 3.
$$ \left\{\begin{array}{l}{\mu }_{i+1}={\mu }_{i}+{\eta }_{\mu }{\sigma }_{i}\cdot {\nabla }_{\mu }J\\ {\sigma }_{i+1}={\sigma }_{i}exp\left(\dfrac{{\eta }_{\sigma }}{2}{\nabla }_{\sigma }J\right)\end{array}\right. $$ (3) $ {\eta }_{\mu } $ and $ {\eta }_{\sigma } $ are the learning rates for μ and σ, respectively. In this study, we set $ {\eta }_{\mu }=1 $ and $ {\eta }_{\sigma }=0.08 $. The segment number is 32×17 and the segment size is 25×50, which means that a range of 800×850 pixels on the DMD is used for light modulation. The light distribution on DMD was calibrated by sequentially turning on each segment and measuring the difference in the output power. In each segment, the number of encoded pixels is 5, and the coding strategy is as previously reported50. It is encoded in the x-axis and expanded to the size of a segment by repeating each pixel in the x- and y-axis (see Section S4, Supplementary information). The population size is 40 and the iteration is 200 for invasive focusing and 400 for non-invasive focusing. These parameters are chosen to balance the optimization speed and the enhancement (see Section S12, Supplementary information). The DMD display speed is mainly limited by the fluorescent signal intensity. 1 kHz is used in the invasive configuration and 200~300 Hz is used in the non-invasive configuration.
-
The UCNP-loaded hydrogel was sonicated at 40 °C for 1 min before it was filled into the rectangular capillary (20 μm × 200 μm, CM Scientific). The capillary was fixed onto the diffuser, which was pasted on a glass slide in order to be clamped by the sample holder. The distance between the holographic diffuser and the resin is 1 mm in air and 14 μm of capillary glass wall; the distance between the chicken tissue and the resin is 170 μm of the coverslip and 14 μm of the glass wall. A “white” pattern was displayed on the DMD, making it just as a mirror. The sample was aligned in the x, y, and z direction so that the capillary is illuminated by the speckle and the speckle size within the capillary is about 30~50 μm. Then the SPAD was aligned to maximize the fluorescent signal.
During the printing, the focusing process was conducted at low power (~7 mW before MO1), and then the focal spot was scanned at higher power (~25 mW before MO2) in order to surpass the polymerization threshold. The hatching distance is 1 μm and the scanning speed is 0.1 μm/s.
UCNPs exhibit stable optical response during 1-hour illumination of 976 nm light (Fig. S14), indicating their reliable performance during printing.
-
A piece of fresh chicken breast was cut into 4-mm cubes and fixed with 10% buffered formalin (HT501128-4L, Sigma Aldrich) overnight. Then the fixed samples were rinsed in phosphate-buffered saline three times and embedded in 2% agarose until solidified. The embedded tissues were cut into 300-μm-thick slices using a vibratome (VT1200 S, Leica), and mounted onto glass slides (Sigma-Aldrich) with Fluoromount-G (SouthernBiotech). 300-μm spacers were used to confine specimens without compression. The sections were sealed with nail polish and kept at 4 °C for 24 hours before being used for printing.
-
This project has received funding from the Swiss National Science Foundation under project number 196971 - “Light based Volumetric printing in scattering resins.” The authors thank the open-source tools (and their contributors) that were used in this work, including Inkscape.org, Python.org, and Fiji.sc. The authors would like to acknowledge Dr. Jessica Sordet-Dessimoz (Histology Core Facility, EPFL) for assistance with tissue sectioning. The authors would like to acknowledge Dr. David Fernando Reyes Vasquez (Interdisciplinary Centre for Electron Microscopy, EPFL) for assistance with transmission electron microscopy. The authors also acknowledge Dr. Yann Lavanchy (Molecular and Hybrid Materials Characterization Center, EPFL) for assistance with FTIR and Zeta-potential characterization.
Light focusing and additive manufacturing through highly scattering media using upconversion nanoparticles
- Light: Advanced Manufacturing , Article number: (2025)
- Received: 11 April 2024
- Revised: 13 January 2025
- Accepted: 13 February 2025 Published online: 03 April 2025
doi: https://doi.org/10.37188/lam.2025.021
Abstract: Light-based additive manufacturing holds great potential in the field of bioprinting due to its exceptional spatial resolution, enabling the reconstruction of intricate tissue structures. However, printing through biological tissues is severely limited due to the strong optical scattering within the tissues. The propagation of light is scrambled to form random speckle patterns, making it impossible to print features at the diffraction-limited size with conventional printing approaches. The poor tissue penetration depth of ultra-violet or blue light, which is commonly used to trigger photopolymerization, further limits the fabrication of high cell-density tissue constructs. Recently, several strategies based on wavefront shaping have been developed to manipulate the light and refocus it inside scattering media to a diffraction-limited spot. In this study, we present a high-resolution additive manufacturing technique using upconversion nanoparticles and a wavefront shaping method that does not require measurement from an invasive detector, i.e., it is a non-invasive technique. Upconversion nanoparticles convert near-infrared light to ultraviolet and visible light. The ultraviolet light serves as a light source for photopolymerization and the visible light as a guide star for digital light shaping. The incident light pattern is manipulated using the feedback information of the guide star to focus light through the tissue. In this way, we experimentally demonstrate that near-infrared light can be non-invasively focused through a strongly scattering medium. By exploiting the optical memory effect, we further demonstrate micro-meter resolution additive manufacturing through highly scattering media such as a 300-μm-thick chicken breast. This study provides a concept of high-resolution additive manufacturing through turbid media with potential application in tissue engineering.
Research Summary
Bioprinting: Overcoming light scattering for printing through biological tissues
Light-based 3D printing offers unmatched precision for creating fine structures. However, printing through biological tissues is severely limited due to strong optical scattering, which prevents light from being focused to a tiny spot and makes it difficult to print at high resolution. Researchers from EPFL have developed a novel method using nanoparticles and light modulating to address this challenge. This technique uses near-infrared light that penetrates deeper through the tissue to excite the nanoparticles, which emit shorter wavelength light to crosslink the structure. Before entering the tissue, light is also shaped by a projector to compensate for the scattering afterward. This technique allows for micro-meter resolution printing through scattering materials like chicken breast tissue.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article′s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article′s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.






 DownLoad:
DownLoad: